Research Areas & Methods
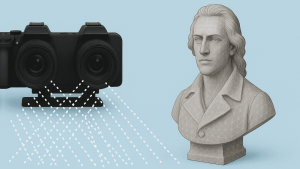
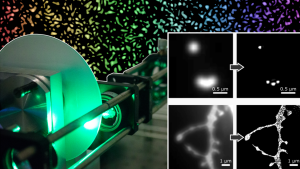
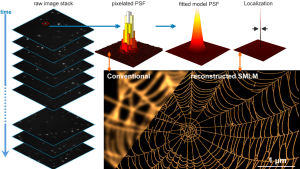
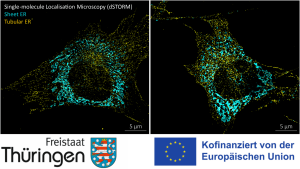
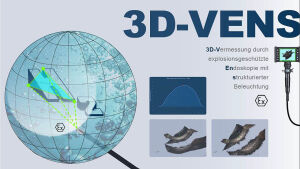
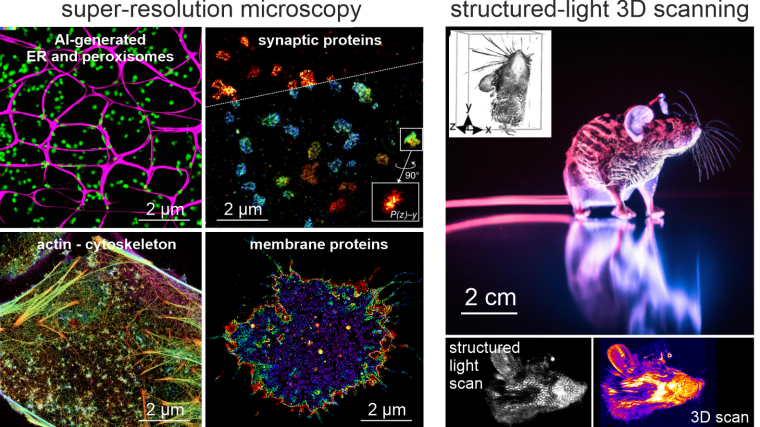
Seeing is believing. The newly established research group of Christian Franke @IAOB de is focused on the development of advanced optical and computational tools for the quantitative study of (cell-) biological questions. On one side, we work to advance super-resolution microscopy methods like single-molecule localization microscopy (SMLM, e.g. dSTORM, PALM) and random structured illumination microscopy (nanoSpeck3D) to quantitatively study the structure-function relationship of sub-cellular (trafficking) organelles, e.g. endosomes, with three-dimensional nanometer resolution. Here, we create novel hard- and software tools to push the limits of 3D volume, colour and time resolution. For this, we have home-built microscopes in our labs in the Abbeanum and are in close collaboration with the Eggeling Group at IAOB and IPHT, providing a large range of complementary microscope setups. Led by Dr. Andreas Stark, we recently branched out into macroscopic measurements of 3D volumes by stereophotogrammetry also utilizing structured illumination, with which we can now measure objects on the centimeter to micrometer scale, thus bridging dimensions between the macro- and nanoscopic world.
Recent Research Results (will be updated soon...)
Recent research includes the development of quantitative tools for novel 3D SMLM approaches [1-3], correlative multi-colour SMLM and volumetric electron tomography (superCLEM)[4] and their application to cell-biological questions [3, 4-6]. For example, we could visualize for the first time the molecular escape of mRNA molecules, delivered by lipid nanoparticles from endosomal recycling tubules at the nanometer scale with multi-colour dSTORM [5] and help to understand part of the fine-structure in developing liver tissue [6]. We also found a novel way of performing macroscopic 3D measurements with a monocular, miniaturized structured-illumination system [7].
[1] Franke et al.External link, Nat. Methods 14, 41-44, (2017).
[2] Franke et al.External link, Nat. Methods 15, 990-992 (2018).
[3] Franke et al.External link, Commun. Biol. 5, 218 (2022).
[4] Franke et al.External link, Traffic 20, 601– 617 (2019).
[5] Paramasivam, Franke et al.External link, J. Cell Biol. 221, e202110137 (2022).
[6] Belicova et al.External link, J. Cell Biol. 220, e202103003 (2021).
[7] Stark et al.External link, Light: Advanced Manufacturing 3, Article number: 34(2022).
[8] Weiler et al.External link, bioRxiv 2022.11.04.515161